Metascience Solutions for the Paradox of Evidence-Based Decision Making
Spencer Phillips Hey, PhD, Harvard Medical School, Boston, MA, USA, and Chief Executive Officer, Prism Analytic Technologies, Cambridge, MA, USA; Joël Kuiper, Chief Technology Officer, Prism Analytic Technologies, Cambridge, MA, USA; Cliff Fleck, Head of Communications, Prism Analytic Technologies, Cambridge, MA, USA
It is easy to say that decisions about planning and design for clinical research should be informed by a comprehensive understanding of the existing evidence. Indeed, in the era of evidence-based medicine, this prescription should seem axiomatic. Whether we are referring to clinical development teams in the pharmaceutical industry planning a translational research program, research funding organizations deciding which project applications to fund, or academic investigators planning and designing their next study, the more that each of these groups understand about what has already been done, what worked well (or did not) in the past, and where the highest value opportunities are, the better.
But while it is easy to say that these kinds of decisions should be based on a comprehensive understanding of the evidence, it is far harder to say exactly how this should be accomplished. In much of the trial methodology literature, for example, it is suggested that these decision makers should refer to the latest, relevant systematic review.1,2 Or if there is no such review, then they should first conduct a systematic review to establish (a) that their proposed line of research has not already been done by someone else, and (b) that their research design decisions are aligned with the best standards and practices.
Unfortunately, conducting a gold-standard systematic review is a laborious and time-consuming process, often taking teams of experts more than 12 months to complete.3 As informative and valuable as this exercise might be, some research and development activities simply cannot be delayed a year or more. Or even if there is already a published systematic review to draw on, given the time it took the authors of that review to complete the work and get it through the peer-review and publication process, the underlying data are likely to be a year or more out-of-date.
Thus, we arrive at one of the paradoxes of evidence-based decision making in clinical research and development: there is widespread agreement that better research will result from first having a comprehensive understanding of the existing evidence in hand. But because time is precious and decisions about what studies to conduct and how to conduct them may not be able to wait a year, a comprehensive, up-to-date understanding is often out of reach.
However, in what follows, we argue that this paradox can be resolved by rethinking some of the fundamental assumptions about the goals and products of systematic evidence reviews.
The Silver Standards of Evidence Synthesis
But before describing our more radical solution, we should first acknowledge that there are a number of alternatives to the gold-standard systematic review.4 Rapid reviews and scoping reviews are 2 methodologies that aim to strike a more time-sensitive balance between the thoroughness of an evidence assessment and gaining sufficient insight to act. The rapid review, for example, will often use a similar search methodology to a full systematic review but will not go as deep into the data extraction or evidence-quality assessment. Or a scoping review may abandon an evidence assessment entirely, and instead focuses solely on providing insight about the size and breadth of the existing evidence base. Both of these approaches can be completed much more quickly than a full systematic review, sometimes taking only days or weeks rather than months. If a primary goal of the evidence review is something like a gap analysis (ie, identifying important questions or clinical needs that have not yet been addressed by any existing studies) then these leaner approaches may be viable alternatives.
It is also worth noting that there are data and analytics companies that aggregate and track data on clinical research programs, or consulting firms that will conduct a complete systematic review and produce recommendations about optimal next steps. For decision makers that have the resources, these companies may be an ideal solution. Data providers may be able to offer all the evidence needed to inform next steps straight “off the shelf,” so to speak. Although consultant firms often still conduct systematic reviews the slow, traditional way, with enough lead time they may provide the richness and depth of analysis needed without requiring their clients to have “personally” expended the time.
However, there are some devils in the details of even these “silver-standard” evidence synthesis options. For example, in the case of purchasing data from a provider, the provenance of the data, as well as the various operations or transformations that have been performed on the data, are not necessarily transparent. This can make it impossible to verify the reliability of the data or any insights derived therefrom. Indeed, this is one of the major reasons for systematic review reporting standards: By documenting every step of the review process, the consumer of the systematic review can have greater confidence that any decisions flowing from it are grounded in a scientifically valid and reproducible process.5
But an even more fundamental challenge for all of these evidence synthesis options is the fact that many of the key terms needed to classify and make sense of biomedical research data are fuzzy, change over time, or are disputed by the experts in the field. For example, even a question as seemingly basic as “What specific disease was being studied in a given clinical trial?” may not have one, clear answer. This could be because the way diseases are classified has changed over time. It could be because there are multiple existing biomedical ontologies, and they do not all perfectly overlap.6 It could also be because the “correct” (or most useful) way to classify the disease of interest in a study depends upon the needs of the decision maker.
This problem of classification is by no means limited to just disease terms. Classifications for drug mechanisms of action, intervention types, population characteristics, inclusion/exclusion criteria, and outcomes are also often fuzzy, disputed, subjective, or context-sensitive. It is therefore necessary for the evidence reviewers to commit to some taxonomy or ontology for these classifiers. For the formal evidence review methodologies, the reviewers are supposed to prespecify the taxonomy/ontology they will use and publish a codebook along with the results that allows the reader to understand their judgment process. Data-providing companies would also ideally make their chosen ontologies and judgment processes explicit for the same reason.
Yet because the problem here is not merely due to multiple classification ontologies but rather stems from fundamental uncertainties in biomedicine (ie, our best understanding is constantly evolving as we learn more), then merely making the ontology and the supporting judgments explicit still does not quite solve it. To truly solve the problem we need to give decision makers a way to quickly aggregate and view relevant evidence with a taxonomy that is suited to their purpose. In other words, the taxonomy of an evidence review also needs to be flexible, so that different “consumers” of the evidence can apply different sets of concepts or classification schemes. Traditional systematic review methods provide all the depth and flexibility needed to do this (because each project can construct its own taxonomy and codebook to suit its purpose), but they are simply too slow. Leaner review methods have speed and flexibility, but may lack the depth needed to optimally inform next steps. The existing “off-the-shelf” data providers have the speed and (often) the depth required, but not the flexibility.
Towards a New Metascience Solution
So can we have it all? Is there a way to give decision makers the speed, depth, and flexibility to evaluate the existing evidence and make the most informed decision possible about what to do next? We believe that the answer to this question is yes but it requires some significant shifts in how we think about the goals and products of an evidence review.
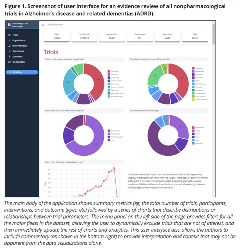
The first shift is to think of an evidence review as a continuous process that should function more like a “living application” that decision makers can interact with and monitor on a regular basis, rather than a one-off, linear project whose end-state will be a static publication. For example, we are currently working on an evidence synthesis project whose goal is to inform pilot trials with promising, nonpharmacological interventions that can improve the quality of care for people with Alzheimer’s disease and related dementias (ADRD).7 To help achieve this goal, we have created an automated search and data extraction algorithm that can regularly query ClinicalTrials.gov for all registered clinical trials and outcomes studies involving nonpharmacological interventions, import these data into a relational database, and then represent the data with a dynamic, visual user interface like the one shown in Figure 1. This approach offers the necessary speed to provide meaningful evidence on the evidence in real-time, without having to wait weeks or months to get an understanding of the scientific landscape before making a decision.
Figure 1. Screenshot of user interface for an evidence review of all nonpharmacological trials in Alzheimer’s disease and related dementias (ADRD).
A second shift is to recognize that all the data transformation (eg, cleaning, normalization, and classification) processes in an evidence review should be transparent. For published systematic reviews, the readers typically only get to see the PRISMA8 flow chart, the summary results (presented in static tables), and perhaps a supplemental spreadsheet with the underlying data in its final state. But the vast majority of judgments that transformed the raw data (which often started as just a list of potentially relevant PubMed/Embase/ClinicalTrials.gov IDs) into a spreadsheet full of valuable classifications remains opaque. Commercial providers are not typically better here since they often do not disclose sufficient details about the processes and algorithms they use. While there may be good business reasons for this opacity, the lack of transparency is anathema to scientific integrity and poses a threat to trust and confidence in the quality of their data.
A third shift, closely related to the second, is to think of the data analysis and presentation as dynamic and evolving. Essentially, we believe that evidence reviews should always be considered ongoing works-in-progress. The data and their analysis will be changing and updating over time, and this is as it should be, given that the scientific community’s understanding is also growing and changing. For example, Figure 2 shows a detailed view of the data-filtering menu from the same ADRD evidence-review user interface mentioned above. This menu allows the user to select/deselect particular types of outcomes and include/exclude those trial records from the analysis. But this particular list of outcome terms, which is the result of a semisupervised, natural language-processing algorithm, is not fixed. As we alter our taxonomy and revise or improve the algorithms over time, this menu and the results of the analysis will change. But since our source data, with their provenance and history of changes, can all be saved in the underlying database, this evolution in how the evidence is processed and interpreted will no longer be problematic. As the science and evidence evolve, so too should our analysis.
Figure 2. Screenshot of filter menu detail for outcome term classifications used for a review of trials in Alzheimer’s disease and related dementias.

In fact, we would argue that the ability to reclassify data and track the history of such changes represents a profound advance in science (or meta-science to be more precise). Indeed, we see this as a key step for overcoming the problems with shifting, inconsistent, or disputed ontologies. Two experts may want to classify the same data in different ways. Each may be justified in their choice of classification and each classification may be correct, given the expert’s particular goal or use-case. But once we come to think of evidence synthesis as an ongoing process and data classification as a dynamic component of this process, we have now transformed this limitation of traditional evidence reviews into a strength.
Conclusion
In sum, the value of evidence-based decision making to guide future research is uncontroversial. Everyone largely agrees that the more we can marshal the “evidence on the evidence,” the more likely we are to make good decisions about next steps and prevent wasteful research. The longstanding challenge has thus been more of a technical one—how can we gather the evidence on the evidence with sufficient speed, reliability, depth, and flexibility? We have argued that these technical challenges can be solved once we stop thinking of evidence synthesis as a linear process that should be done before we act, but rather as an ongoing process of building “living” software applications that give us real-time visibility over the scientific landscape. •
References
1. Chalmers I, Bracken MB, Djulbegovic B, et al. How to increase value and reduce waste when research priorities are set. Lancet. 2014;383(9912):156-165 doi: 10.1016/S0140-6736(13)62229-1.
2. Ioannidis JP, Greenland S, Hlatky MA, et al. Increasing value and reducing waste in research design, conduct, and analysis. Lancet. 2014;383(9912):166-175 doi: 10.1016/S0140-6736(13)62227-8.
3. Ganann R, Ciliska D, Thomas H. Expediting systematic reviews: methods and implications of rapid reviews. Implement Sci. 2010;5(1):56. doi: 10.1186/1748-5908-5-56.
4. Grant MJ, Booth A. A typology of reviews: an analysis of 14 review types and associated methodologies. Health Info Libr J. 2009;26(2):91-108. doi: 10.1111/j.1471-1842.2009.00848.x.
5. Liberati A, Altman DG, Tetzlaff J, et al. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaboration. PLoS Med. 2009;6(7):e1000100. doi:10.1371/journal.pmed.1000100.
6. Faria D, Pesquita C, Mott I, Martins C, Couto FM, Cruz IF. Tackling the challenges of matching biomedical ontologies. J Biomed Semantics. 2018;9(1):4. doi:10.1186/s13326-017-0170-9.
7. NIA Impact Collaboratory. https://impactcollaboratory.org/ Accessed January 12, 2020.
8. Moher D, Liberati A, Tetzlaff J, Altman DG, Prisma Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med. 2009;6(7):e1000097.
Spencer Hey was supported by the National Institute on Aging (NIA) of the National Institutes of Health under Award Number U54AG063546, which funds NIA Imbedded Pragmatic Alzheimer’s Disease and AD-Related Dementias Clinical Trials Collaboratory (NIA IMPACT Collaboratory). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.

