Can We Demonstrate the Value of Next-Generation Sequencing Approaches Within Traditional Value Frameworks?
Bettina Zimmermann, MA, Health System Strategy Leader, F. Hoffmann-La Roche AG, Basel, Switzerland; Daryl S. Spinner, PhD, MBA, Variantyx Inc, Framingham, MA, USA; Susan R. Snyder, PhD, MBA, Georgia State University, School of Public Health, Atlanta, GA, USA; Molly Purser, PhD, MBA, formerly RTI Health Solutions, Research Triangle Park, NC, USA
Introduction
During Virtual ISPOR Europe in Milan, a forum was convened that looked at the question, “Can we demonstrate the value of next-generation diagnostic testing approaches within traditional value frameworks?” which included next-generation sequencing-based testing approaches from reimbursement, ecosystem, and public health perspectives.
The current course of healthcare is unsustainable (Figure),1-4 and according to the World Health Organization (WHO) and the Organization for Economic Cooperation and Development, 20%-40% of healthcare resources are wasted.5
Figure. Unsustainable course of healthcare.
The transformation of healthcare towards precision medicine—giving the right patient the right treatment at the right time6—has the potential to put healthcare on a more sustainable path. Emerging data suggest that multigene next-generation sequencing (NGS)-based testing approaches enable more efficient use of healthcare resources through testing, treatment, and reduction in hospitalization.7-9 When it comes to precision medicine, there is currently a lack of clarity and consensus on a global level on how to assess the value of the technologies and services of precision medicine. In this article, there will be a focus on the challenges of NGS testing approaches as a core element of precision medicine, using mainly US data because most published information is currently available there.
The ISPOR Forum addressed the broader context of next-generation testing (NGT) approaches, which is used as an umbrella term, and covers testing approaches such as multigene NGS, comprehensive genomic profiling, and others (see Table for select examples of multigene NGS tests). Oncology is one of the most advanced medical fields in which molecularly guided treatment options have repeatedly demonstrated delivering better outcomes for patients, and multigene NGS plays an important role in identifying the right patient for the right treatment.11 The number of cancer drugs that need biomarker testing has increased continuously since the late 1990s. This trend is ongoing, so increasingly research and development (R&D) is investigating tumor-agnostic biomarkers (mutations that appear in tumors across indications). Currently, there are over 75 tumor-agnostic treatments in R&D pipelines.12
Table. Examples of multigene NGS tests utilized in current clinical practice.
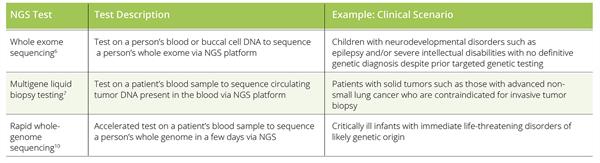
Emerging data suggest that multigene NGS approaches enable more efficient use of healthcare resources through testing, treatment, and reduction in hospitalization.8,9,13 This is important in the context of the current unsustainable course of healthcare spending and existing budget constraints (see Figure).
From a physician’s perspective, the value of NGS-based testing lies in the information that it provides for making quicker and better diagnostic and treatment decisions (ie, which treatments to use or to NOT use, because of resistant mutations3 and clinical trial options, etc).4 Molecular tumor boards have proven to play an important role in interpretation of information as well as enabling efficient healthcare delivery for patients to be treated with options for which there is enough evidence available.3
From a precision public health perspective, the focus is on using data to improve health and achieve social justice—equity, social inclusion, and empowerment. The value of NGS-based testing and similar approaches from a population health perspective is giving the right intervention to the right population at the right time, which is consistent with supporting an individualized treatment approach. Therefore, it contributes to improving the health of identified populations, even if at first glance individualized medicine and public health seem to be in conflict.14 From the outset there were 2 camps: (1) public health and precision medicine present a false dichotomy, since improving population health should include healthcare and molecular tools to stratify populations into risk groups to provide more efficient and effective prevention and treatment strategies, and (2) a more traditional public health view of whole population health that is determined by far more than healthcare. In conclusion, population health outcomes reflect the combined influences of multiple biological and social health determinants, public health efforts and medical care, making “both zip codes and genetic codes” important for health.15
The current course of healthcare is unsustainable and according to the World Health Organization and the Organization for Economic Cooperation and Development, 20%-40% of healthcare resources are wasted.
The core concern of the traditional public health view is whether healthcare system investments in precision medicine’s targeted clinical interventions would come at the expense of existing public health measures that could have a greater impact on population health, and their need for resources. As paradoxical as it seems, while precision medicine focuses on individualized care, its success truly requires a population-based approach.
Value Framework for NGS Testing
Value frameworks, whether utilized explicitly or implicitly in access decisions, have traditionally focused on the payer perspective and are used to assess the value of clinical outcomes of new therapies relative to current therapies. They should help to better inform decisions.16 This assessment sometimes involves evaluating the combination of a test and treatment. Several challenges have been raised in assessing the value of diagnostic approaches, including NGS, when limited to these traditional elements, particularly as the results of multigene NGS testing often provide an end to a prolonged diagnostic odyssey, which may potentially derive from (1) identifying or ruling out one or more potential treatment approaches/further clinical intervention, and/or (2) achieving a definitive diagnosis or ruling out one or more suspected diagnoses. Broadly, the utility of all such information provided by multigene NGS and other testing approaches may be referred to as the “value of knowing,” a healthcare concept introduced decades ago in the literature but formalized more recently in the HEOR field.17,18 Yet when applying a standardized framework, it is important to first consider who the decision maker is, since value may be defined and assessed differently, depending on perspective.
There exist a variety of different value frameworks; some are more centered around medical evidence (eg, National Comprehensive Cancer Network, American Society of Clinical Oncology, European Society for Medical Oncology); others are payer-centered (eg, Institute for Clinical and Economic Review, National Institute for Health and Care Excellence); and some are specific to precision medicine (eg, Personalized Medicine Coalition, MedTech Europe, Genomic Medicine Integrative Research Framework). There is also an ISPOR Value Framework that expands the elements of value beyond traditional cost-effectiveness by incorporating novel value elements like the value of knowing (diagnostic odyssey), real option value (opportunity to benefit from future medical advances), scientific spillover (benefits beyond the original innovation), etc.19
The challenges raised in assessing the value of NGS-based testing are broad, which lead to uncertainty concerning which evidence (endpoints, appropriate and novel trial designs, real-world evidence and emerging applications evolving over time) needs to be made available for patients to be able to access multigene NGS testing and benefit from the interdependent technologies to achieve better outcomes.
Value frameworks provide guidance for evaluating the value of NGS testing, but the current reality is that similar evidence leads to divergent decisions. In the United States, the most recent data from the Centers for Medicare & Medicaid Services (CMS) on Medicare payments show that a sizeable fraction of prescribed NGS testing for Medicare patients is denied reimbursement and that denial rates vary significantly among the payers managing these benefits in their designated regions of the country.2 Decisions on which NGS testing is covered also vary by Medicare payers, contributing to differences in reimbursement and access to such testing across the United States.20
From a precision public health perspective, the focus is on using data to improve health and achieve social justice—equity, social inclusion, and empowerment.
Variability in non-Medicare reimbursement coverage decisions among the multiple national and regional payers also contributes to delays in, or complete barriers to, patient access to NGS testing and potentially to best-informed treatment.21-24 Examples of NGS tests with coverage policies that are divergent across large national and regional payers—despite citing very similar evidence sources in their decision making—include whole exome sequencing in neurodevelopmental disorders, and multigene liquid biopsy testing in oncology (payers reviewed by the authors included Aetna, Anthem BlueCross BlueShield of California, BlueCross BlueShield of North Carolina, Cigna, Humana, and United Healthcare 2020).
A similar situation of divergent NGS test access decisions can be observed across the globe.
Additional uncertainty for prescribers, providers, and developers of NGS-based tests derives from:
• Incentives and priorities of multiple stakeholders
• Nontransparent decision-making criteria/different approaches to valuation
• Limited consideration of unique components of value associated with simultaneously testing multiple analytes
• Nonvalue-based payment with little or no intellectual property protection and high utilization management
• Demand for infeasible drug trial-type study designs
The current access situation calls for the collaboration of all NGS testing stakeholders to create a common ecosystem value of solutions in the context of precision medicine, covering value criteria used for deciding access/adoption, stakeholders to whom the integrated ecosystem value can be proven, and funding/coverage to provide access to clinically useful NGS testing.
Conclusion
There is a need for the current value framework approach for NGS testing access decision making to evolve to a more transparent and harmonized evidence-based approach with clear value criteria addressing the perspectives of multidisciplinary stakeholders covering the entire healthcare ecosystem. This would enable an integrated value-based pathway forward for all stakeholders, most importantly patients, to be able to benefit from these healthcare innovations.
To further clarify the current situation and propose solutions for global NGS testing stakeholders to improve value assessment, 2 ISPOR Special Interest Groups (SIGs)—the Medical Devices and Diagnostics SIG and the Precision Medicine and Advanced Therapies SIG—announced a recently approved joint SIG special project to advance the transparency and evidence basis upon which multigene NGS tests and NGT approaches overall are evaluated and access decisions are made. This work will involve a detailed analysis and consideration of the value assessment landscape along with recommendations aimed at addressing the unique characteristics of NGT as an important innovation towards personalized medicine.
References:
1. United Nations World Population Prospects Highlights. https://population.un.org/wpp/Publications/Files/WPP2019_Highlights.pdf. Accessed August 2020.
2. United Nations World Population Ageing Highlights. https://www.un.org/en/development/desa/population/publications/pdf/ageing/WorldPopulationAgeing2019-Highlights.pdf. Accessed August 2020.
3. The Cancer Atlas. The Burden of Cancer. https://canceratlas.cancer.org/the-burden/the-burden-of-cancer/ Accessed August 2020.
4. Deloitte Global Healthcare Outlook. https://www2.deloitte.com/content/dam/Deloitte/global/Documents/Life-Sciences-Health-Care/gx-lshc-hc-outlook-2019.pdf. Accessed August 2020.
5. World Health Organization. The World Health Report. Health Systems Financing: The Path to Universal Coverage. http://digicollection.org/hss/documents/s18314en/s18314en.pdf. Accessed June 1,2020.
6. Srivastava S, Love-Nichols J-A, Dies KA, et al. Meta-analysis and multidisciplinary consensus statement: exome sequencing is a first-tier clinical diagnostic test for individuals with neurodevelopmental disorders. Genet Med. 2019;21(11):2413-2421.
7. Colomer R, Mondejar R, Romero-Laorden N, Alfranca A, Sanchez-Madrid F, Quintela-Fandino M. When should we order a next generation sequencing test in a patient with cancer? EClinicalMedicine. 2020;25:100487. doi: 10.1016/j.eclinm.2020.100487.
8. Pennell NA, Mutebi A, Zhou ZY, et al. Economic impact of next-generation sequencing versus single-gene testing to detect genomic alterations in metastatic non–small-cell lung cancer using a decision analytic model. JCO Precision Oncology. 2019;3:1-9.
9. Yencho S, Austin J, Betka E. Cancer healthcare utilization impact of precision therapeutics: hospitalization/emergency visits. Cancer Sci Res. 2020:3(2):1-3.
10. Sweeney NM, Nahas SA, Chowdhury S. Rapid whole genome sequencing impacts care and resource utilization in infants with congenital heart disease. NPJ Genom Med. 2021;6(1):29.
11. Subbiah V, Kurzrock R. Challenging standard-of-care paradigms in the precision oncology era. Trends Cancer. 2018;4(2):101-109.
12. The IQVIA Institute. IQVIA.com. Supporting precision oncology: targeted therapies, immuno-oncology, and predictive biomarker-based medicines. Accessed February 9, 2021.
13. Coquerelle S, Darlington M, Michel M, et al. Impact of next generation sequencing on clinical practice in oncology in France: better genetic profiles for patients improve access to experimental treatments. Value Health. 2020;23(7):898-906. doi: 10.1016/j.jval.2020.03.005.
14. Horton R. Offline: in defence of precision public health. Lancet. 2018;392(10157):1504. doi: 10.1016/S0140-6736(18)32741-7.
15. CDC Genomics & Precision Health Blog. https://www.cdc.gov/genomics/blog/index.htm#anchor_1499859659149. Accessed August 1, 2020.
16. Neumann P J, Willke RJ, Garrison, LP Jr. A health economics approach to US value assessment frameworks—introduction: an ISPOR Special Task Force report [1]. Value Health. 2018;21(2):119-123. doi: 10.1016/j.jval.2017.12.012.
17. Garrison LP, Jr, Kamal-Bahl S, Towse A. Toward a broader concept of value: identifying and defining elements for an expanded cost-effectiveness analysis. Value Health. 2017;20(2):213-216. doi: 10.1016/j.jval.2016.12.005.
18. Takashima S, Nomura N, Noguchi Y, Matsuzuka F, Inoue T. Primary thyroid lymphoma: evaluation with US, CT, and MRI.
J Comput Assist Tomogr. 1995;19(2):282-288.
19. Lakdawalla DN, Doshi JA, Garrison LP, Jr, Phelps CE, Basu A, Danzon PM. Defining elements of value in health care—a health economics approach: an ISPOR special task force report [3]. Value Health. 2018;21(2):131-139. doi: 10.1016/j.jval.2017.12.007.
20. Centers for Medicare & Medicaid Services. cms.gov. Medicare Provider Utilization and Payment Data. https://www.cms.gov/Research-Statistics-Data-and-Systems/Statistics-Trends-and-Reports/Medicare-Provider-Charge-Data. Accessed September 10, 2020.
21. Hsiao SJ, Sireci AN, Pendrick D, et al. Clinical utilization, utility, and reimbursement for expanded genomic panel testing in adult oncology. JCO Precision Oncology. 2020;4:1038-1048.
22. Park SK, Thigpen J, Lee IJ. Coverage of pharmacogenetic tests by private health insurance companies. J Am Pharm Assoc. 2003;60(2):352-356.e353. doi: 10.1016/j.japh.2019.10.003.
23. Reuter CM, Kohler JN, Bonner D, et al. Yield of whole exome sequencing in undiagnosed patients facing insurance coverage barriers to genetic testing. J Genet Couns. 2019;28(6):1107-1118.
24. Sireci AN, Aggarwal VS, Turk AT, Gindin T, Mansukhani MM, Hsiao SJ. Clinical genomic profiling of a diverse array of oncology specimens at a large academic cancer center: identification of targetable variants and experience with reimbursement. J Mol Diagn. 2017;19(2):277-287.
.


